| COOKIES: By using this website you agree that we can place Google Analytics Cookies on your device for performance monitoring. | ![[Talks.cam]](https://talks.cam.ac.uk/images/talkslogosmall.gif?1209136071) |
University of Cambridge > Talks.cam > Biophysical Seminars > Structural Analysis of Amyloid-like Protein Fibrillation
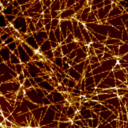
Structural Analysis of Amyloid-like Protein FibrillationAdd to your list(s) Download to your calendar using vCal
If you have a question about this talk, please contact Jerome Charmet. All welcome Protein amyloid fibrillation is associated with a number of grave diseases, most notably the neurodegenerative diseases such as Alzheimer’s and Parkinson’s diseases (1). Protein and peptide fibrillation also constitutes a major challenge in the biopharmaceutical industry, where fibrillation must be avoided to ensure product safety (2). The structural investigation of protein fibrillation is however inherently challenging, since a number of structural species co-exist during the fibrillation reaction. These species cover a wide range of sizes (nm to µm) and exist in different volume fractions over time, in an equilibrium that is highly sensitive to the experimental conditions. Isolation of individual species is thus not possible. At the same time, it is important to investigate the structural species formed during the fibrillation pathway. Not only are these key to understanding the molecular principles behind the process, but also accumulating evidence links such intermediate species to cytotoxic activity, central to the progressive, degenerative diseases. We use small angle X-ray scattering (SAXS) as a central method to investigate the fibrillation reaction. Formation of -synuclein (aSN) fibrils is associated with Parkinson’s disease. We have previously characterized the low-resolution structure of intermediately formed aSN oligomers (3) and reveal that these oligomers are building blocks of the fibril structure (4). We have recently demonstrated, that early amyloidogenic aSN species can disrupt lipid model systems, and that lipid:protein co-aggregates in a non-amyloid state are formed in this context (5) while the effect on lipid membranes varies depending on the lipid composition (6). While the methodology behind the data analysis from such complex systems has been well elaborated both by us (7), and others (e.g. 8), the need for a robust, objective and (semi-)automated analysis system is evident, and our latest efforts in this direction will be presented. We apply the newly developed software to the analysis of a familial mutant of aSN, revealing the occurrence of intermediate species which have a different nature than those previously characterized (unpublished results). References: 1. Eisenberg & Jucker (2012) Cell, 148, 1188-1203 2. Das (2012) AAPS PharmSciTech 13, 732-746 3. Giehm et al. (2011) PNAS , 108, 3246-3251 4. Pedersen et al. (2015) Scientific Reports, 5, 10422 5. van Maarschalkerweerd et al (2014) Biomacromolecules 15, 3643-3654 6. van Maarschalkerweerd et al. in review 7. Langkilde & Vestergaard (2012) Methods. Mol. Biol. 849, 137-155 8. Lorenzen et al. (2014) JACS 136 , 3859-3868 This talk is part of the Biophysical Seminars series. This talk is included in these lists:
Note that ex-directory lists are not shown. |
Other listsCyber Security Society talks SBR Graduate Talks home Workshop on Multimodal Approaches to Language Acquisition Millennium Maths Project public and schools' events How representing multiple objects (and features) as an ensemble enhances higher-level visual cognitionOther talksThe persistence and transience of memory Richard Horton (The Lancet Cheif Editor): Scientific Publishing 5 selfish reasons to work reproducibly Finding meaning in English writing Designer Babies or Children of Frankenstein? Genome Editing and its Side Effects Participatory approaches to encourage responsible use of antibiotics in livestock |