| COOKIES: By using this website you agree that we can place Google Analytics Cookies on your device for performance monitoring. | ![[Talks.cam]](https://talks.cam.ac.uk/images/talkslogosmall.gif?1209136071) |
University of Cambridge > Talks.cam > Biophysical Seminars > The properties of nano-scale amyloid fibril fragments in vitro and in vivo
The properties of nano-scale amyloid fibril fragments in vitro and in vivoAdd to your list(s) Download to your calendar using vCal
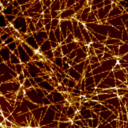
If you have a question about this talk, please contact Jerome Charmet. All welcome What are the molecular mechanisms that govern the amyloid fibrils’ potential to seed the formation of new amyloid to facilitate their spreading, and to damage cells in amyloid-associated diseases such as Alzheimer’s disease, Parkinson’s disease, type 2 diabetes, and systemic amyloidoses? These disease-associated properties of amyloid have been linked to small nano-sized particles created through the fragmentation of amyloid fibrils, which is a process governed by the stability of amyloid fibrils toward breakage. We have previously described an approach that is capable of resolving fibril fragmentation rates, fibril particle concentrations, and fibril length distributions, through direct observations by atomic force microscopy image analysis. Applying our approach to several different amyloid model systems, such as alpha-synuclein, lysozyme, and the yeast prion protein Sup35NM, we present an unique comparison of the stability of these different amyloid fibrils toward breakage. Using the yeast prion protein Sup35NM that forms amyloid particles capable of conferring district yeast phenotypes in vivo, we have also quantified the biological impact of fibril fragments in terms of their transmissive potential. Our results allow new quantitative insights into key properties of amyloid aggregated in terms of their their destructive association to disease as well as the constructive potential they have for future functional amyloid materials development. This talk is part of the Biophysical Seminars series. This talk is included in these lists:
Note that ex-directory lists are not shown. |
Other listsCambridge International Development Conference 2015 Caius Medsoc Talks: "A Timeline of Medicine" Physics of Medicine (PoM) Seminar Series facisnating Talks Seminars at the Department of Biochemistry British Society of Aesthetics Cambridge Lecture SeriesOther talksPlanck Stars: theory and observations Far-infrared emission from AGN and why this changes everything Poison trials, panaceas and proof: debates about testing and testimony in early modern European medicine Future directions panel "Itsa me! Luigi!" [citation needed] - unlocking your referencing skills Aspects of adaptive Galerkin FE for stochastic direct and inverse problems |